A high throughput platform for the rapid characterization of glycosylation in recombinant proteins, serum proteins, exosomes, or ECM proteins
The technology utilizes a panel of nearly 50 different lectins as probes for testing the presence of a large number of biologically relevant glycan binding motifs and structural elements simultaneously on N- and O-glycans as well as glycolipids without requiring previous deglycosylation.
We have over 15 years of experience in lectin microarray development and analysis covering recombinant glycoproteins, serum glycoproteins, synthetic neoglycoproteins, cells, exosomes, or extracellular matrix (ECM) proteins.
Over 48 highly purified lectins isolated from natural sources with well-established glycan binding specificities are printed in 5 replicates alongside control samples (printing buffer containing BSA). We print our lectin microarrays on Premium Schott Nexterion 3D hydrogel glass slides with an ultralow background and excellent protein stabilization properties.
If you have larger sample numbers or are interested in a specific lectin panel, please inquire about custom array development.
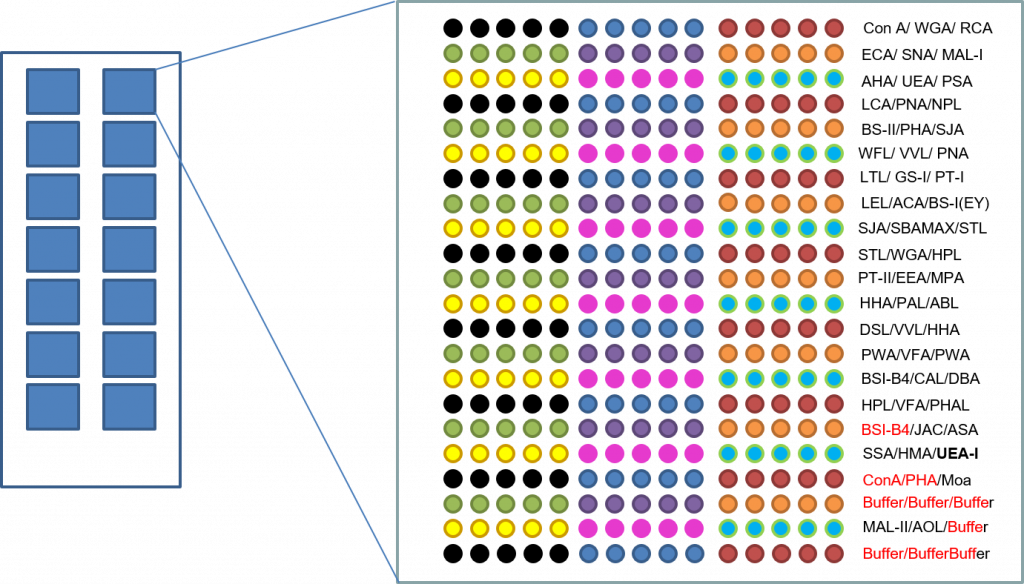
Method Summary
Glycoanalysis is performed in triplicate.
Glycoprotein samples are fluorescently labeled, and incubated on the lectin array, and the interactions are quantified by fluorescence detection. Semiquantitative binding data is provided as histograms showing the corresponding lectin interaction profiles for each sample. Raw data (xlsx format) and a fluorescence image of the incubated lectin microarrays are also provided.
| Type | Lectin | Specificity | Monosaccharide |
|---|---|---|---|
| AAA | Anguilla anguilla lectin | α-L-fucose | α-L-fucose |
| AAL | Aleuria Aurantia | Fuc | Fuc |
| ABL | Agaricus bisporus | Gal(β1-3)GalNAc, T antigen | Gal |
| ACA | Amarantus Caudatus | T antigen, (Galβ1-3GalNAcα-Thr/Ser) | Gal |
| AHP | Arachis hypogaea AHA | Gal(β1-3)GalNAc, T Antigen | Gal |
| AIA/JAC | Artocarpus integrifolia (Jacalin) | T antigen | GalNAc |
| AOL | Aspergillus orzyae | Fucose | Fuc |
| ASA | Allium sativum lectin | α1,3-mannose | Man |
| BPL | Bauhinia Purpurea | Galβ-1,3 or β1,4-GalNAc | Gal |
| BS-I/GS-I | Griffonia simplicifolia I | α-Gal | Gal |
| BS-II/GS-II | Griffonia simplicifolia II | Terminal GlcNAc | GlcNAc |
| CAL | Cicer arietinum | Fetuin, Lac, IgM | Lac |
| CFL | Codium fragile | GalNAc | GalNAc |
| ConA | Concanavalin A | α-Man, α-Glc | Man |
| DBL/DBA | Dolichos biflorus | α-GalNAc | GalNAc |
| DSL | Datura stramonium | (GlcNAc)2, LacNAc | GalNAc |
| ECA | Erythrina Cristagalli A | Gal, GalNac | GalNAc |
| EEA | Euonymus Europaeus | Lac, blood groups B and H | Gal |
| GNA | Galanthus Nivalis | terminal α1,-3Man | Man |
| HAL | Helix aspersa | GalNAc | GalNAc |
| HMA | Homarus americanus | LAg1: NeuNAc. LAg2: GalNAc | Sialic acid, GalNAc |
| HPL | Helix pomatia | α-GalNAc terminal | GalNAc |
| LBA | Phaseolus lunatus | GalNAc-α1,3[Fuc-α1,2]Gal | GalNac |
| LEL | Lycopersicon esculentum | (GlcNAc)3 | GlcNAc |
| LTL | Lotus tetragonolobus | Terminal α-Fuc, Lex | Fuc |
| MAL-I | Maackia amurensis Lectin I | α2,3-SialLacNac | Sial |
| MAL-II | Maackia amurensis Lectin II | LacNAc | Gal |
| MOA | Marasmium oreades agglutinin | Gal-α1,3-Gal and Gal-α1,3-Gal-β1,4-GlcNAc | α-Gal |
| MPA | Maclura Pomifera | T antigen, α-GalNAc | Gal |
| NPL | Narcissus Pseudonarcissus | Terminal and internal Man | Man |
| PAL | Pseudomonas aeruginosa PA-I | Gal | Gal |
| PHA E+L | Phaseolus Vulgaris Agglutining | oligomers | Lac |
| PNA | Peanut agglutinin | T antigen, Gal(β-1,3) GalNac | Gal |
| PSA | Pisum sativum | Fucα-1,6-GlcNAc and α-Man | Fuc |
| PT-I | Psophocarpus tetragonolobus I | α-GalNAc | GalNAc |
| PT-II | Psophocarpus tetragonolobus II | α-1,2-fucosylated LacNAc | Gal |
| PWA | Phytolacca americana | (GlcNAc)3 | GlcNAc |
| RCA120 | Ricinus Communis Agglutinin, | β-Gal, Lac, LacNAc | Gal |
| SBA | Soybean agglutinin | α-Gal-GalNAc | GalNAc |
| SJA | Sophora japonica | GalNAc | GalNAc |
| SNA | Sambucus nigra | α-2,6 sialic acid on LacNAc | Sialic, Lac |
| SSA | Salvia sclarea lectin | Terminal GalNAc linked to serine | GalNAc |
| STL | Solanum tuberosum | (GlcNAc)3, LacNAc | GlcNAc |
| UEA-I | Ulex Europaea Aggl | α-1,3-Fuc. L-fucose | Fuc |
| VFA | Vicia faba lectin | α-Man, Glc, GlcNAc | Mannose |
| VVL | Vicia villosa B4 | GalNAc | GalNAc |
| WFL | Wisteria floribunda | GalNAc | GalNAc |
| WGA | Triticum Vulgaris | (GlcNAc)n, sialic acid | GlcNAc |
Promotional offer
Our current offer for lectin array analysis :
- Optional sample preparation and protein isolation step (not included)
- Sample labeling with fluorescent dye and sample cleanup
- Optimization of sample concentration and incubation conditions on test slide
- Lectin binding analysis in triplicate (4 samples/slide)
- Microarray quantification, data analysis, and data visualization as histograms
- Report including histograms, data file, and discussion
Price upon request
Please inquire about customized quotations for larger sample numbers or if you do not require triplicate measurements.

